Project name: NOPP-gigas-ploidy-temp
Funding source: National Oceanographic Partnership Program
Species: Crassostrea gigas
variable: ploidy, elevated seawater temperature, desiccation
Background:
RNA extractions from ctenidia were performed using the lab RNazol protocol. Tissue used was from 72 diploid and triploid oysters:
- sdate: July 12, 2021 - 12 diploid and 12 triploid - controls
- sdate: July 12, 2021 - 12 diploid and 12 triploid - 30C
- sdate: July 13, 2021 - 12 diploid and 12 triploid - 30C + desiccation
The sample list, with RNA results, can be found here.
The concentration of extracted RNA was first estimated using the nanodrop. These values were then used to dilute RNA for analysis on the Agilent 2100 bioanalyzer using RNA 6000 Pico chips, which have a detection range of 50-5000 pg/ul for Total RNA in water.
Here are the results of the first 11 samples that I ran today:
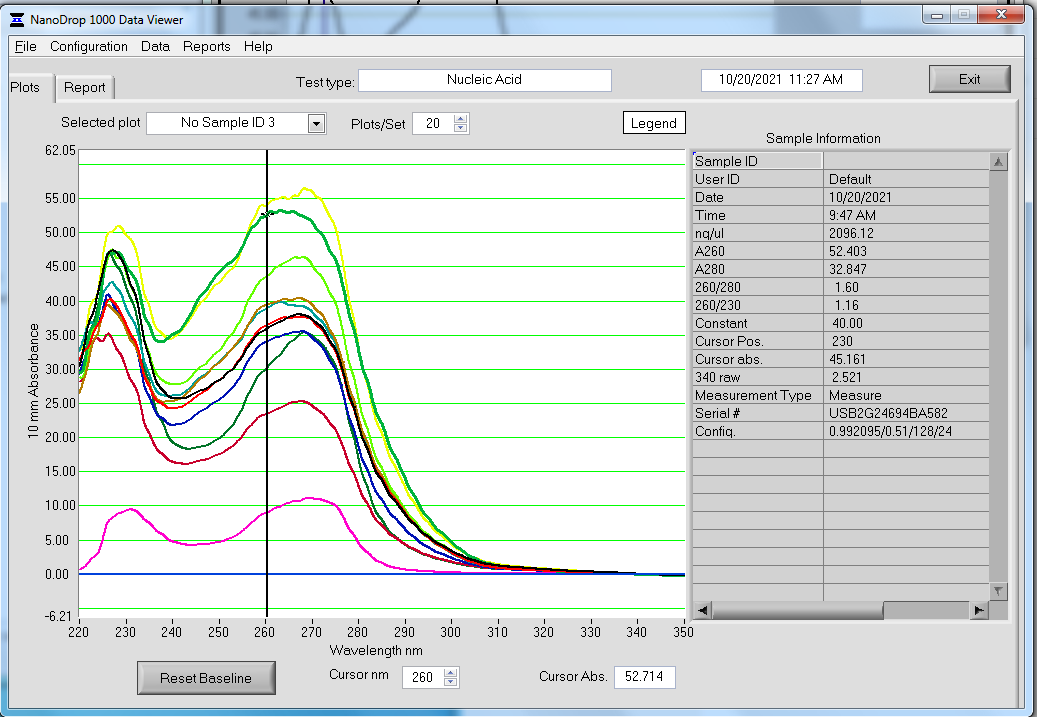
Nanodrop - UV spectra
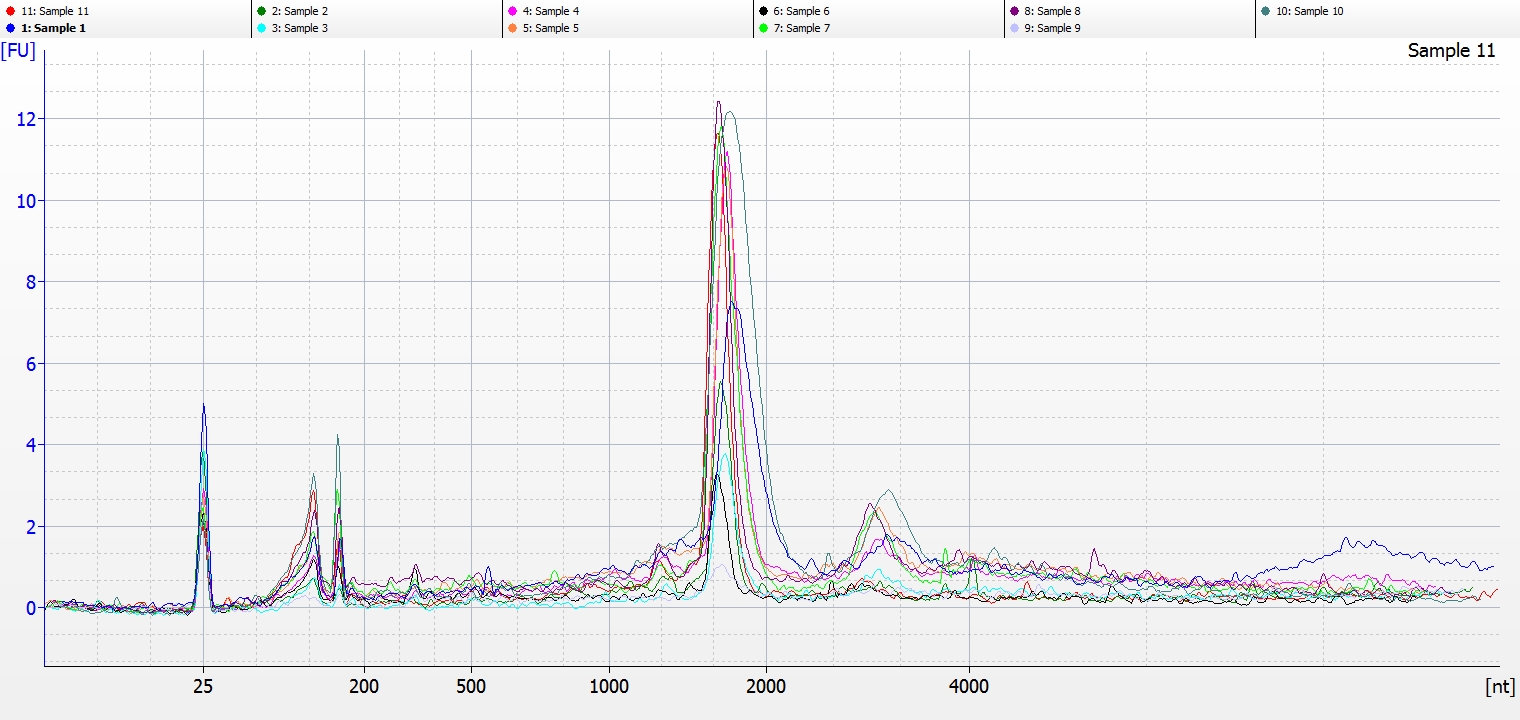
Bioanalyzer - electropherogram (overlay)
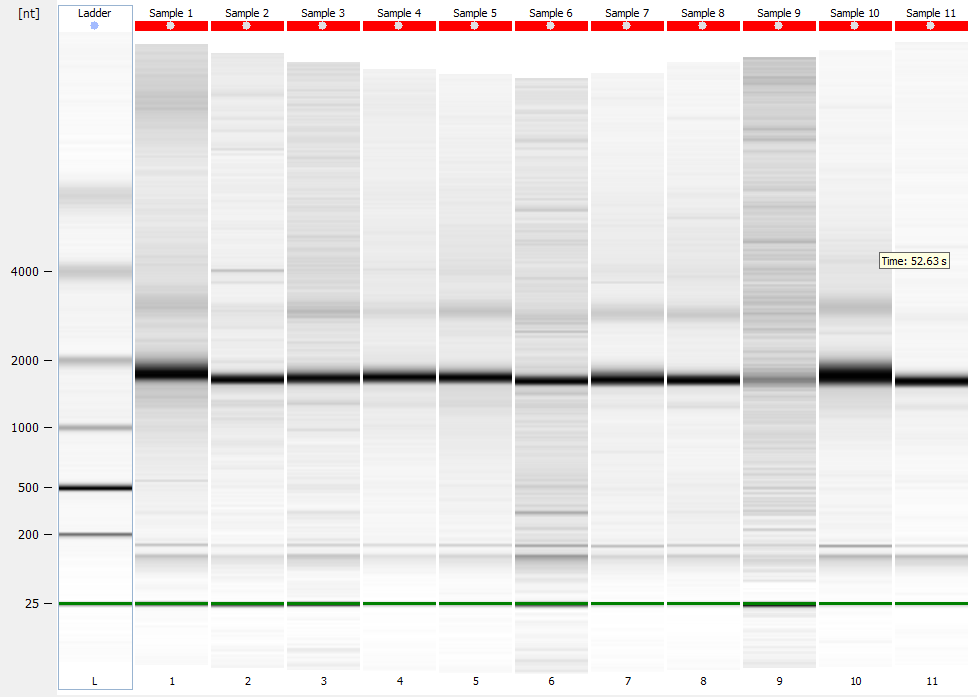
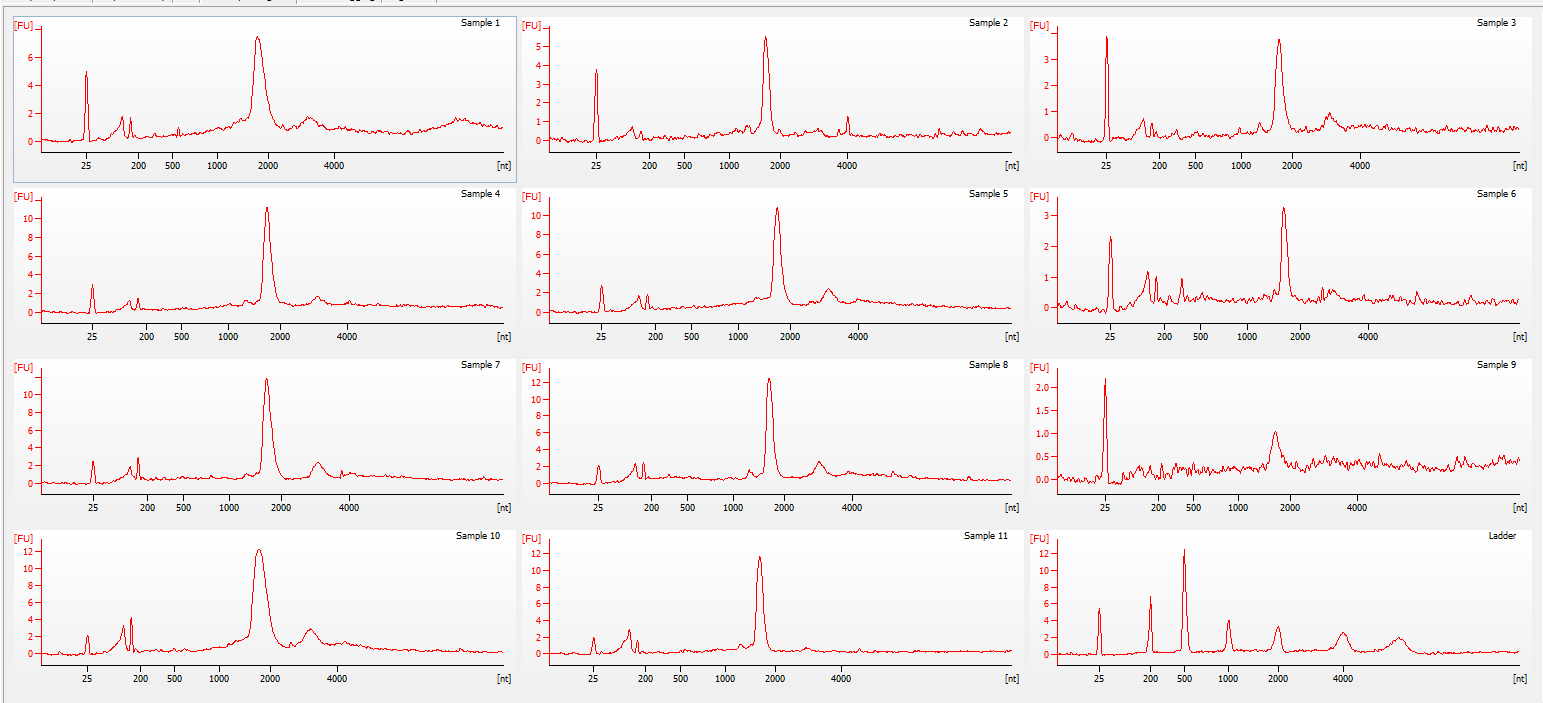
Bioanalyzer - electropherogram (individual)
Bioanalyzer - gels